Currently, Sonarqube is not supported on EKS. There is documentation around how to install it on Docker, however no clear documentation around setting up the cluster on ECS or EKS. So, here you learn how to install Sonarqube?
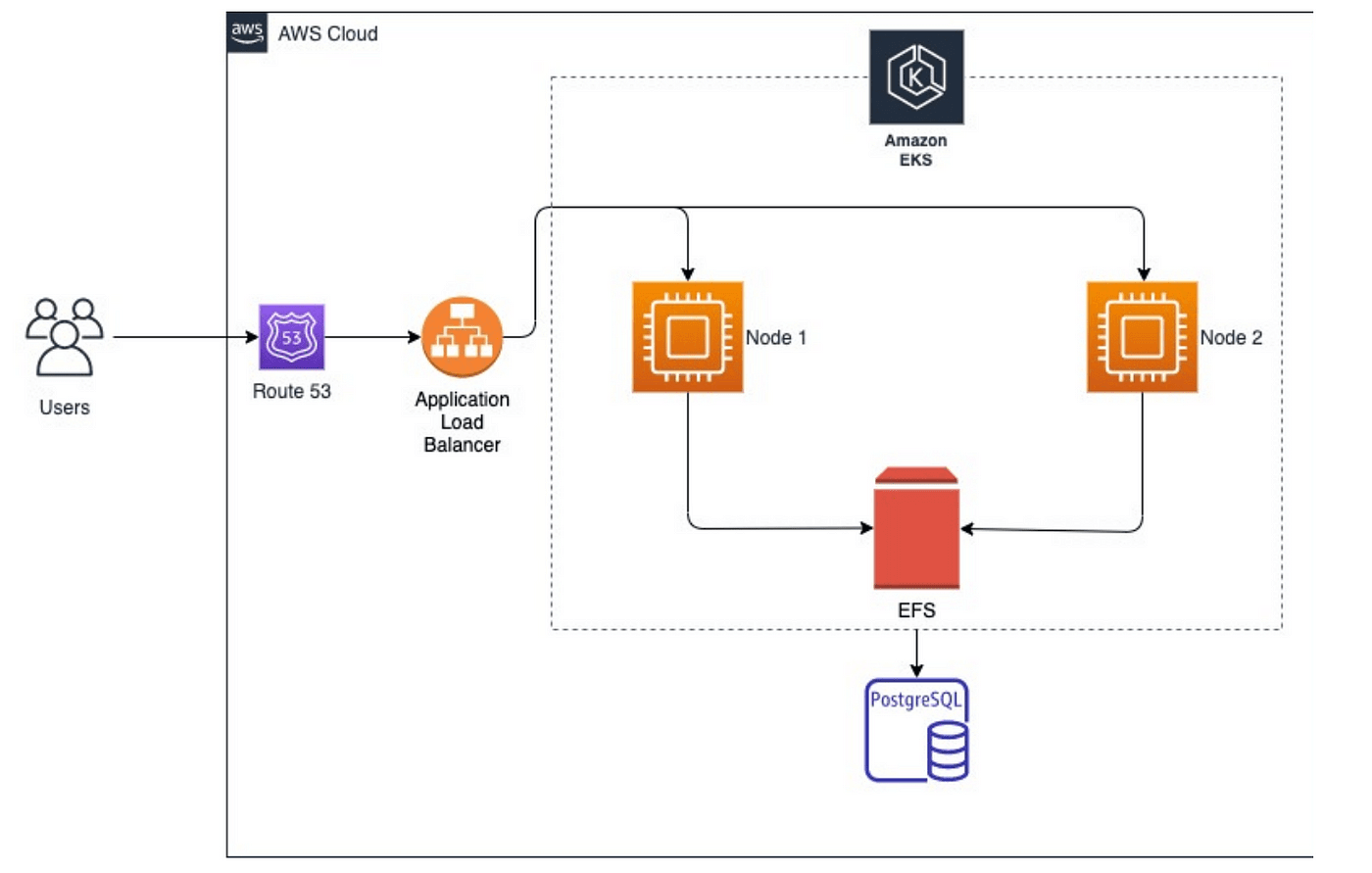
We will try to implement the below architecture where we will create an EKS cluster using the eksctl command and use EFS as persistent storage for data generated from the Sonarqube application.
The article doesn’t take into consideration of security practices . This document is purely for reference purposes.
Create an IAM User with Admin Permissions
Navigate to IAM > Users. Click Add user. Set the following values: User name: k8-admin, Access type: Programmatic access Click Next: Permissions. Select Attach existing policies directly. Select AdministratorAccess. Click Next: Tags > Next: Review. Click Create user. Copy the access key ID and secret access key, and paste them into a text file, as we’ll need them in the next step. Also read: Latest DevSecOps Trends
Launch an EC2 Instance and Configure the Command Line Tools Navigate to EC2 > Instances. Click Launch Instance. On the AMI page, select the Amazon Linux 2 AMI . Leave t2.micro selected, and click Next: Configure Instance Details. On the Configure Instance Details page: Click Next: Add Storage > Next: Add Tags > Next: Configure Security Group. In the key pair dialog, select Create a new key pair. Give it a Key pair name Click Download Key Pair, and then Launch Instances. Click View Instances, and give it a few minutes to enter the running state. Once the instance is fully created, check the checkbox next to it and click Connect at the top of the window. In the Connect to your instance dialog, select EC2 Instance Connect (browser-based SSH connection). Click Connect. In the command line window, check the AWS CLI version:aws --version It should be an older version. Download v2:
curl “https://awscli.amazonaws.com/awscli-exe-linux-x86_64.zip” -o “awscliv2.zip”
See where the current AWS CLI is installed:
It should be /usr/bin/aws Update it:
sudo ./aws/install –bin-dir /usr/bin –install-dir /usr/bin/aws-cli –update
Check the version of AWS CLI:
It should now be updated.
For AWS Access Key ID, paste in the access key ID you copied earlier. For AWS Secret Access Key, paste in the secret access key you copied earlier. For the Default region name, enter us-west-2 For Default output format, enter json. Download kubectl:
curl -o kubectl https://amazon-eks.s3.us-west-2.amazonaws.com/1.16.8/2020-04-16/bin/linux/amd64/kubectl
Apply execute permissions to the binary:
Copy the binary to a directory in your path:
mkdir -p $HOME/bin && cp ./kubectl $HOME/bin/kubectl && export PATH=$PATH:$HOME/bin
Ensure kubectl is installed:
kubectl version –short –client
curl –silent –location “https://github.com/weaveworks/eksctl/releases/latest/download/eksctl_$(uname -s)_amd64.tar.gz” | tar xz -C /tmp
Move the extracted binary to /usr/bin:
sudo mv /tmp/eksctl /usr/bin
Get the version of eksctl:
See the options with eksctl:
Provision an EKS Cluster Provision an EKS cluster with two worker nodes in us-west-2:
eksctl create cluster –name sonarqube –version 1.21 –region us-west-2 –nodegroup-name sonarqube-workers –node-type t3.medium –nodes 2 –nodes-min 2 –nodes-max 3 –node-volume-size 30 –ssh-access –managed
Update kubeconfig on bastion node
aws eks update-kubeconfig –name sonarqube –region us-west-2
Run kubectl command to check nodes
Provision EFS
Ref: https://docs.aws.amazon.com/eks/latest/userguide/efs-csi.html
Create Postgres instance in EKS using helm
Ref: https://dev.to/arctype/deploy-and-manage-postgresql-on-kubernetes-oni
Create a new database for Sonarqube
postgres=# l
Create a Secret to store PostgreSQL password Kubernetes has a built-in capability to store secrets. To create a secret you need to base64 encode a secret value
echo -n ” | base64
and create a k8s secret using YAML file
apiVersion: v1
Apply the secret file
kubectl apply -f db_password.yml
Create a SonarQube deployment After creating PVCs and Postgres secret we are ready to deploy using the following YAML file
apiVersion: apps/v1
Apply the deployment file
kubectl apply sonarqube-deployment.yml
Publish SonarQube via Kubernetes Service After SonarQube is up and running we need to create a public endpoint to access our service
apiVersion: v1
———-
$ kubectl apply -f sonar-svc.yaml
Now we can hit EXTERNAL-IP address and login to SonarQube
Limitations Only one replica set can be run for Sonarqube LTS or community edition. Multiple replica set is only available on Data-center edition